Differential expression between groups using pseudobulk
Sarah Williams
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6096346/
Overview
Once we have identified cell types present in the samples, its common to test how gene expression changes between experimental conditions, within each different cell type.
Some cell types may be dramatically affected by the experimental conditions, while others are not. Likewise some genes may change only in a specific cell type, whereas others show a more general difference.
This document describes how to apply a pseudobulk approach to test for differences between groups. In a pseudobulk approach counts are obtained by pooling together groups of cells; in this case cells from the of the same celltype from the same sample. These pooled counts can then be analysed more like a bulk RNAseq experiment.
This is very similar to how a non-spatial single cell experiment may be analysed.
Note that there are many other approaches to calculate differential expression in this kind of data - including those that make use of individual cells; see review (Soneson and Robinson 2018).
This test requires:
- Biological replicates for each group
- Assigned cell types (or clusters, or niches)
For example:
- What genes are differentially expressed in epithelial cells in Crohn’s disease vs healthy individuals?
- How do genes change with treatment in each different cell type in my sample?
- What differences are there between treatment sensitive and resistant cancer cell? (instead of a ‘celltype’ could use a cancer or stromal niche)
Steps:
- Calculate pseudobulk
- Filter to testable pseudobulk groups (enough cells to pool)
- Filter to testable genes (enough expression to see changes)
- Test for changes in gene expression
- Plot DE results and individual genes.
Worked example
How does gene expression change within each cell type between Ulcerative colitis or Crohn’s disease, and Healthy controls?
Using data from Macrophage and neutrophil heterogeneity at single-cell spatial resolution in human inflammatory bowel disease from (Garrido-Trigo et al. 2023);
The study included 9 cosmx slides of colonic biopsies
- 3x HC - Healthy controls
- 3x UC - Ulcerative colitis
- 3x CD - Chrones’s disease
Load libraries and data
library(alabaster.sfe)
library(SpatialExperiment)
library(scuttle)
library(scater)
library(tidyverse)
library(limma)
library(DT)
library(edgeR)
library(BiocParallel)
library(Voyager)
library(patchwork)
library(ggrepel) # gg_repel, For non-overlapping gene labelsdata_dir <- file.path("~/projects/spatialsnippets/datasets/GSE234713_IBDcosmx_GarridoTrigo2023/processed_data")
sfe_01_loaded <- file.path(data_dir, "GSE234713_CosMx_IBD_sfe_01_loaded")
se_pseudobulk_file <- file.path(data_dir, "GSE234713_CosMx_IBD_pseudobulk_sample_se2.RDS")sfe <- readObject(sfe_01_loaded)Experimental design
There are three individuals per condition, one tissue sample from each individual. 9 slides in total.
Note: Each tissue sample has multiple FOVs captured. In the cosMX output an ‘fov’ refers to the rectangular regions on the slide that are measured; there are multiple fovs per sample, and in this experiment, only one sample per slide. It may be neccessary to remove ‘poor’ FOVs or sampels with too few FOVs, but not in this experiment.
sample_table <- select(as_tibble(colData(sfe)), condition, individual_code, fov_name) %>%
unique() %>%
group_by(condition, individual_code) %>%
summarise(n_fovs= n(), item = str_c(fov_name, collapse = ", "))
DT::datatable(sample_table)We have 5 broad celltypes defined already.
plot_one <- function(sample) {
p <- plotSpatialFeature(sfe[,sfe$tissue_sample == sample], "celltype_subset",colGeometryName = "cellSeg") +
theme(legend.title=element_blank()) +
ggtitle(sample)
return(p)
}
samples <- levels(sfe$tissue_sample)
plots <- lapply(FUN=plot_one, X=samples)
p.all <- wrap_plots(plots, ncol=3)
plot(p.all)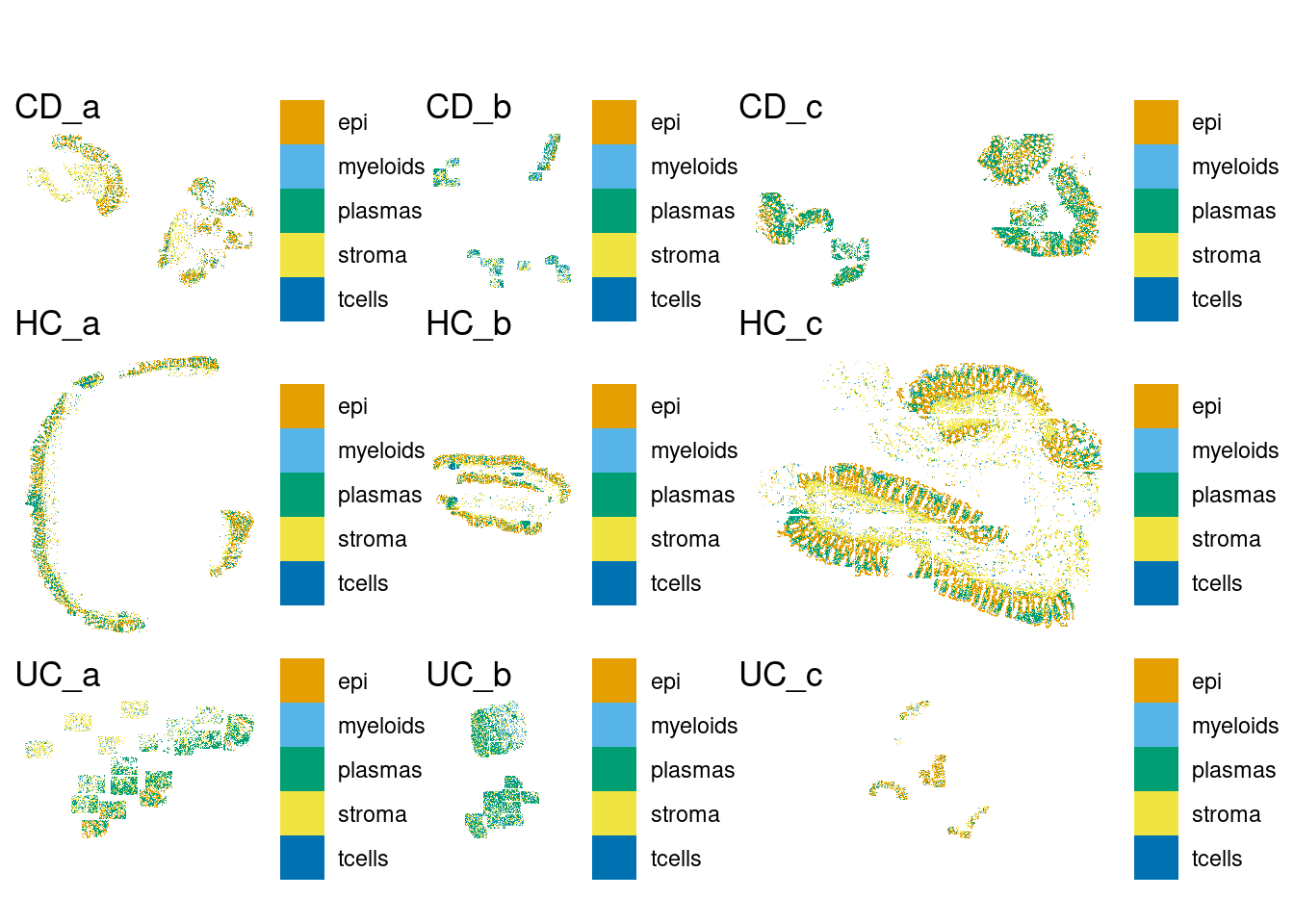
| Version | Author | Date |
|---|---|---|
| 44907e6 | swbioinf | 2025-11-07 |
Cell filtering
We need to define the grouping that will make one pseudobulk sample for analysis. In this experiment, we can consider all cells within a tissue sample as somewhat equivalent. Therefore that is all cells of a particular celltype, within a tissue sample.
If we had tiled fovs across a tissue sample with distinct regions (e.g. cancer, stroma) it might be necessary to group them by subregion as well.
Testing other things
We are not limited to tests at the celltype level between conditions, tests could be at any informative grouping. E.g.
- Differences in expressesion at a tissue niche level (e.g. ) - Define pseudobulk groups as tissue_sample + niche and test between condition.
- Differences in expression of celltype localised within niche vs outside of niche - Design pseudubulk groups as tissue_sample + celltype + niche
This same approach can be used to these hypothesese by adjusting the pseudobulk grouping and limma design model (further below) accordingly. This example assumes tests between conditions at the celltype level.
Once the pseudobulk replicate grouping is decided, we need to check there will be enough cells to perform our analyses. Note there are several levels of filtering here!
- Need at least x reads in a cell to include it
- Need at least x cells of a celltype within an fov to include a sample
- Can only test where we have at least 2 samples on each side of a contrast.
- In some datasets, it may be worth removing entire samples that have too few fovs (e.g. Tumour microarrays with some samples peeling off the slide). This isn’t neccesary for this study.
What these thresholds should be needs to be determined for each experiment.
Filter counts per cell
To start, check out the distribution of reads per cell. Here, we choose and apply a filter of 200 reads. This is low compared to what you might see in a single scRNAseq experiment, but we are working with only 1000 genes.
Even if you have alreay applied a counts per cell filter across your dataset (e.g at least 50 counts per cell), you might choose to apply a second, higher, threshold for differnetial expression. We can afford to be more stringent for the differential expression threshold because the spatial information isn’t being used, so there is no problem of ‘gaps’ between cells. Exact thresholds will depend on your data. More counts in a cell tend to result in better cell type identification and smoother expression values and should theoretically yeild better differential expression results - if there are enough cells.
min_reads_per_cell <- 200
ggplot(colData(sfe), aes(x=total_count)) +
geom_density() +
geom_vline(xintercept = min_reads_per_cell, lty=3) +
scale_x_log10() +
theme_bw()+
ggtitle("How many reads per cell?")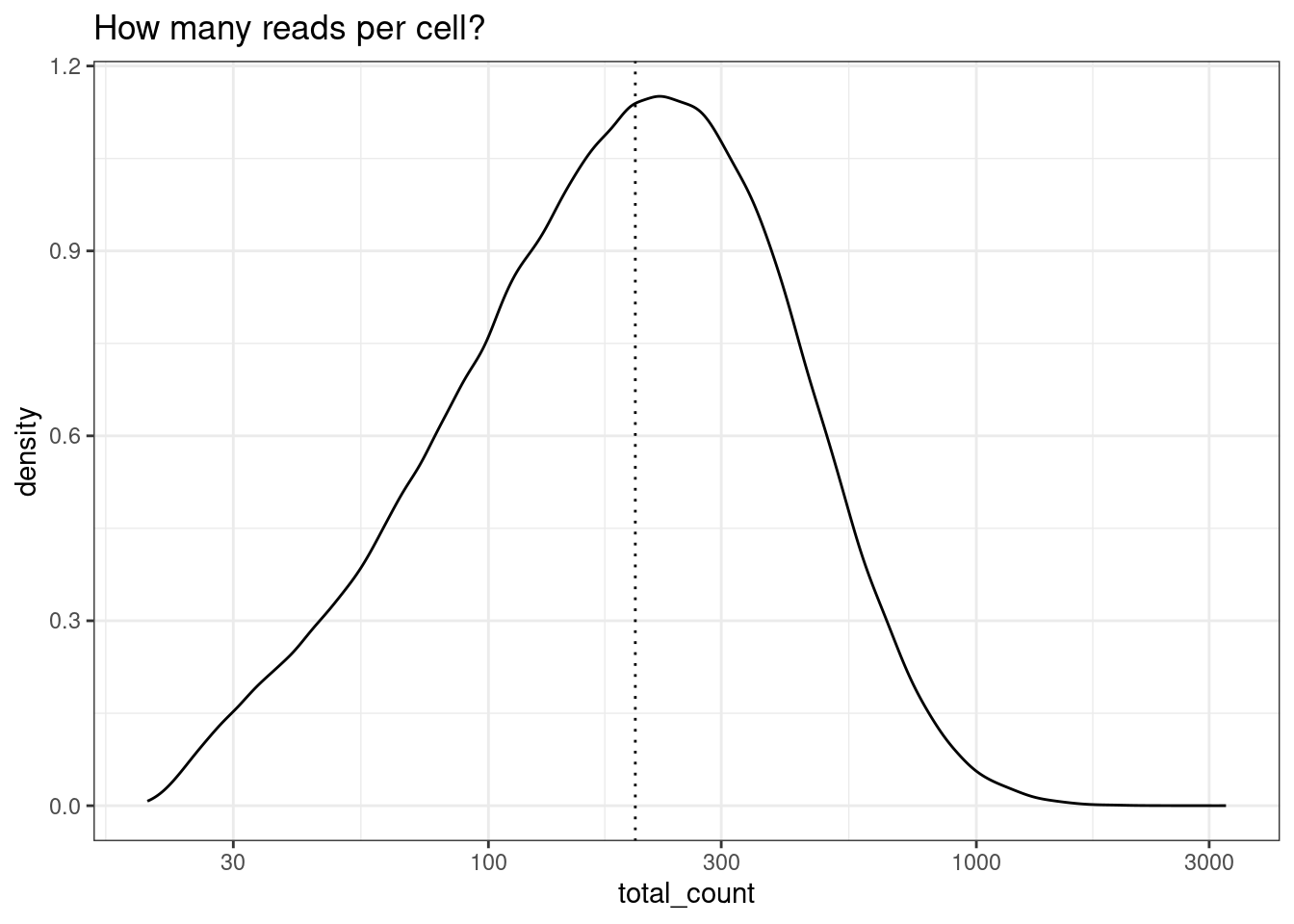
| Version | Author | Date |
|---|---|---|
| 44907e6 | swbioinf | 2025-11-07 |
sfe <- sfe[,sfe$total_count>= min_reads_per_cell]Calculate pseudobulk
We need to created a cell annotation for the pseudobulk grouping in our analysis. We can do this by concatenating the tissue sample ID with the celltype; calling this column pdb_sample here.
Now use the PseudobulkExpression() function to sum up each gene’s gene expression across each pdb_sample.
This will build a SummarisedExperiment object at the ‘sample_cluster’ level’. Which contains one entry per group, containing the sum of copies of the gene in all cells within the grouping.
This can take a while to run - this code uses 8 cores to speed it up. Even so, its well worth saving the ‘pseudobulk’ object to disk for later calculations and visualisations.
sfe$pdb_sample <- paste0(sfe$tissue_sample, '_', sfe$celltype_subset)
se.pdb <- aggregateAcrossCells(sfe, ids=sfe$pdb_sample)
# Use 8 cores, requires BiocParallel,
#se.pdb <- aggregateAcrossCells(sfe, ids=sfe$pdb_sample,
# BPPARAM = MulticoreParam(workers=8) )
# To save it
saveRDS(se.pdb, se_pseudobulk_file)Read in the previously prepared object.
se.pdb <- readRDS( se_pseudobulk_file)Now instead of counts for each individual cell, we have our pseodubulk matrix with the pooled sum of counts for each celltype within each tissue samples. So the numbers are much higher, with fewer zeros.
But note that each pool is a different size, so we can’t compare counts directly. We will address this later with normalisation.
counts(se.pdb)[1:10,1:4] CD_a_epi CD_a_myeloids CD_a_plasmas CD_a_stroma
AATK 306 31 85 206
ABL1 363 15 74 327
ABL2 310 30 76 158
ACE 284 15 38 94
ACE2 508 25 111 176
ACKR1 191 16 66 268
ACKR3 209 16 51 142
ACKR4 313 16 59 127
ACTA2 197 20 38 481
ACTG2 195 27 64 1044Pseudobulk sample filtering
Once the pseudobulk replicate grouping is decided, its time for the next level of filtering. We need to check there will be enough cells within a celltype within the sample to perform our analyses.
Filter cells per group
Each of our ‘pseudobulk’ samples represents a pool of cells. But some pools will contain very few cells, and should be excluded.
The aggregateAcrossCells function we used to group the groups keeps values in the sample annotation where every cell has the same value (or NA if they don’t) - this means we have all the sample and condition information. Plus, it adds a new column called ‘ncells’, which is the number of cells in the pool.
DT::datatable(data.frame(colData(se.pdb))[,c('pdb_sample','individual_code','tissue_sample','ncells')])# What is the minimum accepptable number of cells in a pool?
min_cells_per_pdbsample <- 100
ggplot(colData(se.pdb), aes(x=ncells, col=celltype_subset)) +
geom_density() +
geom_vline(xintercept=min_cells_per_pdbsample, lty=3) +
geom_rug() +
scale_x_log10() +
theme_bw() +
ggtitle("How many cells per pseduobulk sample?")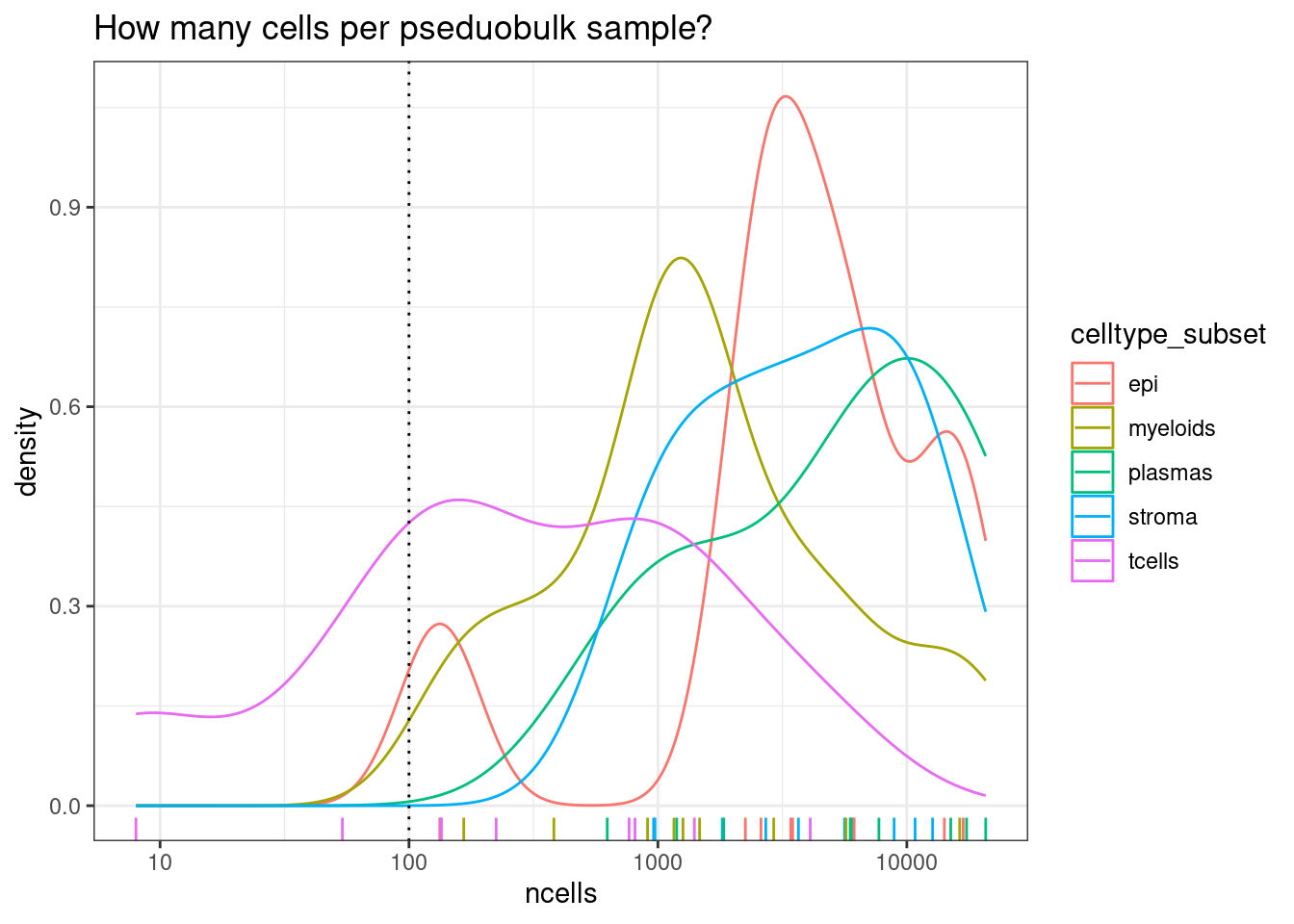
| Version | Author | Date |
|---|---|---|
| 44907e6 | swbioinf | 2025-11-07 |
Note there are much fewer t-cells overall. Its typical that some of the less common cell types are difficult or impossible to reliably test.
We can then remove pools where there were too few cells - choosing a threshold of 100 in this instance.
ncol(se.pdb)[1] 45se.pdb <- se.pdb[,se.pdb$ncells >= min_cells_per_pdbsample]
ncol(se.pdb)[1] 43Samples per contrast
Ignoring pseodureplicate fovs from a single sample, how many biological samples are represented in our filtered table?
In this experiment, there are enough samples to run the contrasts for every cell type.
celltype_summary_table.passed <- data.frame(colData(se.pdb)[,c('pdb_sample','condition','individual_code','celltype_subset')])
celltype_summary_table.passed %>% select(condition, individual_code, celltype_subset) %>%
unique() %>%
group_by(condition, celltype_subset) %>%
summarise(n=n()) %>%
pivot_wider( names_from=condition, values_from = n)# A tibble: 5 × 4
celltype_subset `Healthy controls` `Ulcerative colitis` `Crohn's disease`
<fct> <int> <int> <int>
1 epi 3 3 3
2 myeloids 3 3 3
3 plasmas 3 3 3
4 stroma 3 3 3
5 tcells 2 3 2Calculate Differential Expression
We now have a counts matrix, and an annotation table that describes its samples.
This looks very much like a bulk RNAseq experiment. Except - instead of X samples across Y conditions, we have X samples of Y conditions for each of Z celltypes.
# For clarity, pull out the counts amtrix and annotation table
pseudobulk_counts_matrix <- counts(se.pdb)
pseudobulk_anno_table <- as_tibble(colData(se.pdb)[,c('pdb_sample','group','condition','individual_code','tissue_sample','celltype_subset')])
pseudobulk_counts_matrix[1:10,1:4] CD_a_epi CD_a_myeloids CD_a_plasmas CD_a_stroma
AATK 306 31 85 206
ABL1 363 15 74 327
ABL2 310 30 76 158
ACE 284 15 38 94
ACE2 508 25 111 176
ACKR1 191 16 66 268
ACKR3 209 16 51 142
ACKR4 313 16 59 127
ACTA2 197 20 38 481
ACTG2 195 27 64 1044head(pseudobulk_anno_table)# A tibble: 6 × 6
pdb_sample group condition individual_code tissue_sample celltype_subset
<chr> <fct> <fct> <fct> <fct> <fct>
1 CD_a_epi CD Crohn's dis… CD_a CD_a epi
2 CD_a_myeloids CD Crohn's dis… CD_a CD_a myeloids
3 CD_a_plasmas CD Crohn's dis… CD_a CD_a plasmas
4 CD_a_stroma CD Crohn's dis… CD_a CD_a stroma
5 CD_b_epi CD Crohn's dis… CD_b CD_b epi
6 CD_b_myeloids CD Crohn's dis… CD_b CD_b myeloids We will use a fairly standard limma differential expression analysis.
We will process each celltype one at a time as follows:
- Subset the counts matrix and annotation table to the celltype.
- Build and fit the model. In our case, a simple one that looks for various by disease group, blocked on individual.
- Check there are enough biological samples to run the test
- Run the test.
How to build the model and run the test will need to be customised for your experimental design - it is very flexible to support paired analyses, batches and other factors. Online resources that describe approaches for bulk RNAseq analyses can be applied to pseudobulk analyses - see the ‘More Information’ section of this document for suggestions.
The model
In this case the line model.matrix( ~0 + group) builds a
model where gene expression is expected to vary by group (UC/CD/HC). The
~0 part indicates a ‘intercept’, which is convenient because it means we
can include all 3 groups in our contrasts, rather than one of them being
treated as the baseline. ( NB: Mathematically, you could use ~group
alone without intercept, and define contrasts accordingly to get the
same result. )
Some example models:
- ~0 + group : Most straightforward approach taken here- one pooling fovs tile across a sample, tested in parallel for each celltype.
- ~0 + group, (blocked on individual with using duplicate Correlation) : Generally not needed, takes into account pseodureplicate fovs from the same samples (if we expect some particularly large variation between fovs it may be considered). We do this by blocking on individual, and calculating a intra duplicate correlation with duplicateCorrelation()(Smyth, Michaud, and Scott 2005). That’s a method from microarray times, see discussion for how it is applied to bulk RNAseq data.
- ~0 + group + individual : If this was a treatment / time point experiment with paired samples from individuals. Other experimental factors can be added this way as needed (e.g. slide, phenotype)
- ~0 + group + celltype : Instead of subsetting to celltype, includes cell type in the model. This pulls more data into the statistics, but makes building the contrasts challenging! Not covered here.
About Filtering
What if you do need to skip constrasts? Often its easiest to count your biological replicates at the differential expression step and skip them there. You might need to run contrasts between groups one at a time to do this.
Actually calculate differential expression
# Build a table of each contrast we might want to do.
#contrasts = group A - group B.
#UCvHC = UC - HC
#CDvHC = CD - HC
contrasts_wanted <- bind_cols(
A= c("UC", "CD"), # First terms
B= c("HC", "HC") # Second terms (usually control)
)
# Empty list to collect results
de_result_list <- list()
## Cycle through each celltype
for (the_celltype in levels(se.pdb$celltype_subset)) {
# Subset pseudobulk object to one celltype
se.pdb.this <- se.pdb[,se.pdb$celltype_subset == the_celltype]
# And pull out the annotation and counts
anno_table.this <- as.tibble(colData(se.pdb.this))
count_matrix.this <- counts(se.pdb.this)
## Check for sufficient replicates ##
# To do any calculations, we need at least 2 pseudobulk groups per contrast.
# there are plenty in this experiemnt, but with less replicates and rare cell types
# its very common to have to skip some contrats for some celltypes.
# skip clusters with no samples after filtering
if( nrow(anno_table.this) < 1 ) {next}
# Count how many bio samples per group (only need one fov to count it)
biosample_per_group <- anno_table.this %>%
select(group, tissue_sample) %>%
unique() %>% # Don't care how many (passed) fovs there were on the tissue
pull(group) %>% # pull out group column, one entry per tissue sample
table() # count how many
# Are there enough Biological smples to consider the contrast?
min_biosample_per_group <- 2
enough_biosamples <-
(unname(biosample_per_group[contrasts_wanted$A]) >= min_biosample_per_group) &
(unname(biosample_per_group[contrasts_wanted$B]) >= min_biosample_per_group)
# Make a new table with contrasts that we have enough biosamples for.
contrasts_to_test <- contrasts_wanted[enough_biosamples,]
## Setup model
# Setup objects for limma
dge <- DGEList(count_matrix.this)
dge <- calcNormFactors(dge)
# Build model
group <- anno_table.this$group
individual_code <- anno_table.this$individual_code
# Model design
# Add other experimental factors here
# ~0 + group
# ~0 + group + individual
# ~0 + group + individual + slide
design <- model.matrix( ~0 + group)
# Run Voom
vm <- voom(dge, design = design, plot = FALSE)
# Fit model
fit <- lmFit(vm, design)
# Define and fit contrasts and run ebayes
# Doing this in an automated way so we can include/exvlude contrasts where
# there are / arenot enough replicates available.
# That's why generatgin a string .
contrast_str_list <- paste0("group",contrasts_to_test$A,"-","group",contrasts_to_test$B)
contrasts <- makeContrasts(contrasts=contrast_str_list,
levels=coef(fit))
fit <- contrasts.fit(fit, contrasts)
fit <- eBayes(fit)
## Loop through contrasts.
# You ccan run multiple contrasts at onces, but doing it this way allows us
# to skip individual contrasts within a celltype.
for ( the_coef in colnames(contrasts) ) {
# Make a version of the coeffient name that doesn't include a '-', or the
# factor name (group)
# Life is easier without special characters
# groupUC-groupHC => UCvHC
contrast_name <- gsub("group","", gsub("-","v",the_coef))
de_result.this <- topTable(fit, n = Inf, adjust.method = "BH", coef = the_coef) %>%
rownames_to_column("target") %>%
mutate(contrast=contrast_name,
contrast_group="pairwise",
celltype=the_celltype) %>%
select(celltype,contrast_group, contrast,target,everything()) %>%
arrange(P.Value)
# build a unique name for this result by adding
# <celltype>_<A>v<B>
de_result_list[[paste(the_celltype, contrast_name, sep="_")]] <- de_result.this
}
}
# Join together results for all celltypes, and pull out those with a singificant adjusted p-value
de_results_all <- bind_rows(de_result_list)
de_results_sig <- filter(de_results_all, adj.P.Val < 0.05)Table of significant results.
DT::datatable(mutate(de_results_sig, across(is.numeric, signif, digits = 3)))DE plots
The below plots show the logFC calculate for each gene versus its average expression across all samples. This is a useful diagnostic plot to evaluate your differential expression results.
- Only big changes are significant at lower expressions, simply becuase higher expression means more statistical confidence.
- A lopsided plot might indicate some oddness around the normalisation of your data (e.g. extremely different cell counts). We can see that here, but note the significant genes are those deviating from the trend.
make_ma_style_plot <- function(res_table, pval_threshold = 0.05, n_genes_to_label = 10) {
p <- ggplot(res_table, aes(x=AveExpr, y=logFC, col=adj.P.Val < pval_threshold) ) +
geom_hline(yintercept = c(0), col='grey80') +
geom_point(pch=3) +
geom_text_repel(data = head(arrange(filter(res_table , adj.P.Val < pval_threshold ), P.Value), n=5),
mapping = aes(label=target), col="red" ) +
theme_bw() +
geom_hline(yintercept = c(-1,1), lty=3) +
scale_colour_manual(values = c('FALSE'="black", 'TRUE'="red")) +
theme(legend.position = 'none')
return(p)
}#res_table.UCvHC.epi <- filter(de_results_all, contrast == "UCvHC", celltype=="epi")
p1 <- make_ma_style_plot(res_table = filter(de_results_all, contrast == "UCvHC", celltype=="plasmas")) + ggtitle("UC vs HC - plasmas")
p2 <- make_ma_style_plot(res_table = filter(de_results_all, contrast == "UCvHC", celltype=="tcells"))+ ggtitle("UC vs HC - T-cells")
p3 <- make_ma_style_plot(res_table = filter(de_results_all, contrast == "UCvHC", celltype=="stroma")) + ggtitle("UC vs HC - stroma")
p1 + p2 + p3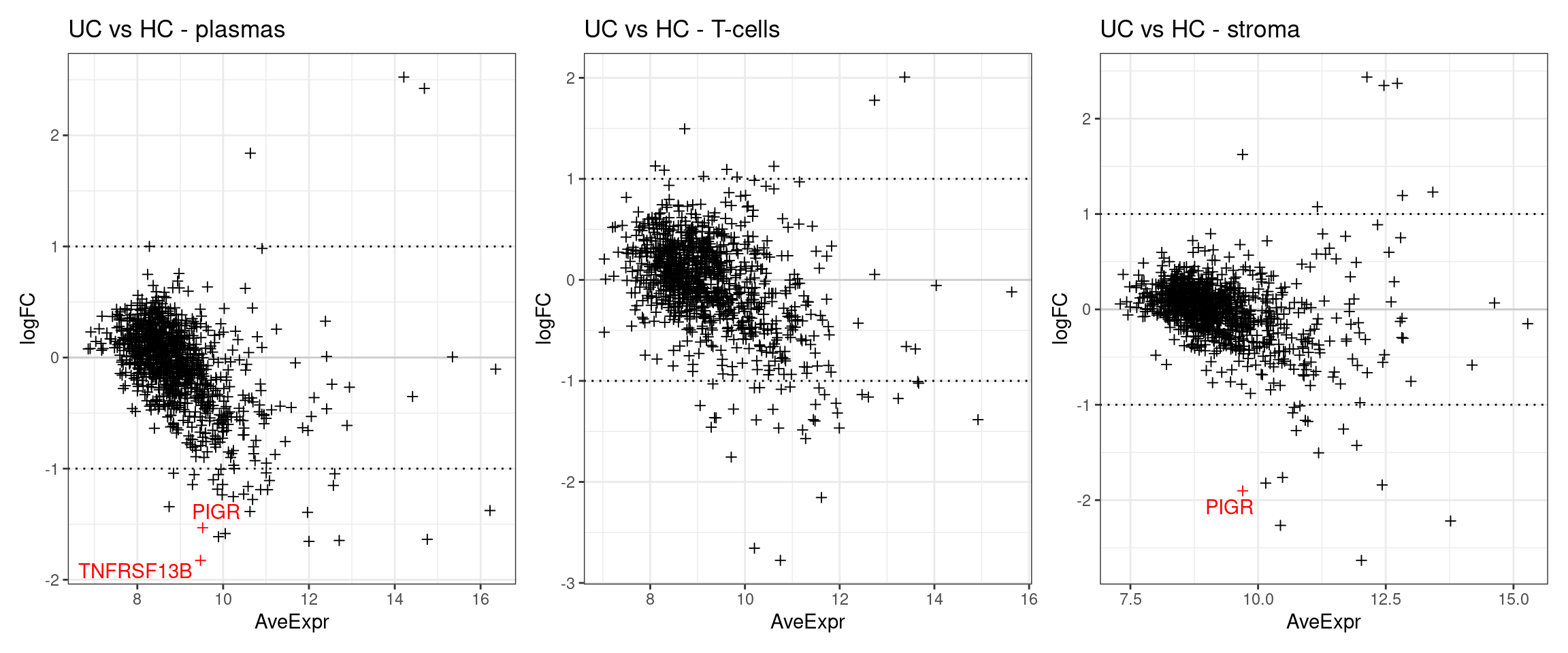
Check some examples
Its always worth visualising how the expression of your differentially expressed genes really looks, with respect to your experimental design. How best to do this depends on your experiment.
The results suggests that LYZ was significantly DE between individuals with Ulcerative Colitis and Healthy Controls in epithelial cells. We can easily plot the log2 normalised pseudobulk scores for each grouping to see how it changes between groups, celltypes and conditions. There’s a fair variability between individuals, not surprising for human data.
# Get some normalised values, purely for plotting.
# (can't use logged counts because the sizes are so dramatically different!)
dge <- DGEList(counts(se.pdb))
dge <- calcNormFactors(dge)
norm_counts <- cpm(dge)
# Store normalised log2 scale expression in
assay(se.pdb, "logcounts") <- log2(norm_counts)DT::datatable(mutate(filter(de_results_sig, target == "LYZ"), across(is.numeric, signif, digits = 3)))plotExpression(se.pdb, "LYZ", x="group", colour_by = "group",
other_fields = "celltype_subset") +
facet_wrap(~celltype_subset)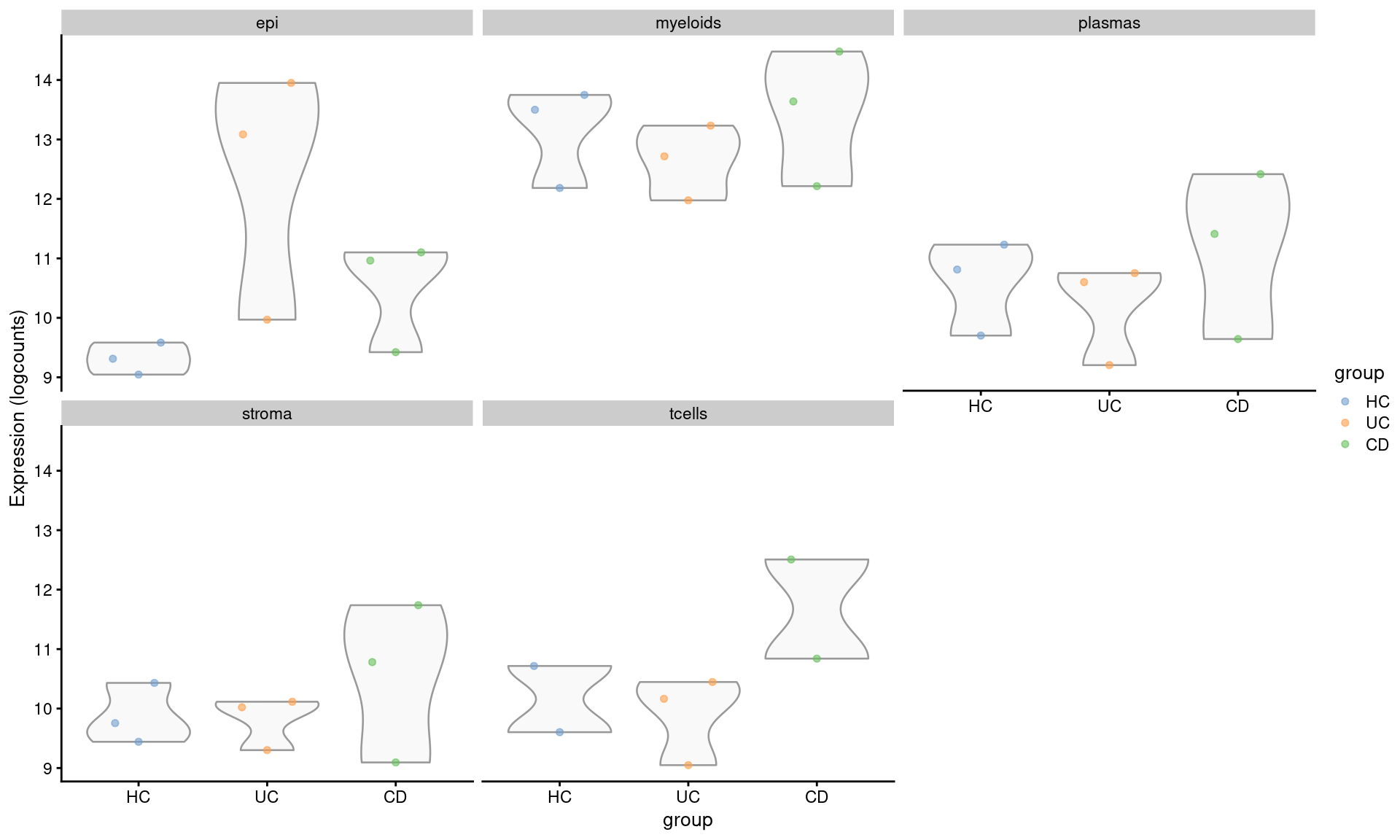
| Version | Author | Date |
|---|---|---|
| 755e4ca | swbioinf | 2025-11-07 |
PIGR was flagged more globally across multiple comparisons, in plasma and stroma in Crohns disease vs healthy, and stroma and myeloids when comparing Ulcerative colitis vs healthy. PIGR is a more specifically expressed at a higher level in the epithelia however - so we would want to interpet this carefully (e.g. Are stroma cells really expressing less PIGR, or could they be catching expression from more-oft co-located epithelia?)
DT::datatable(mutate(filter(de_results_sig, target == "PIGR"), across(is.numeric, signif, digits = 3)))plotExpression(se.pdb, "PIGR", x="group", colour_by = "group",
other_fields = "celltype_subset") +
facet_wrap(~celltype_subset)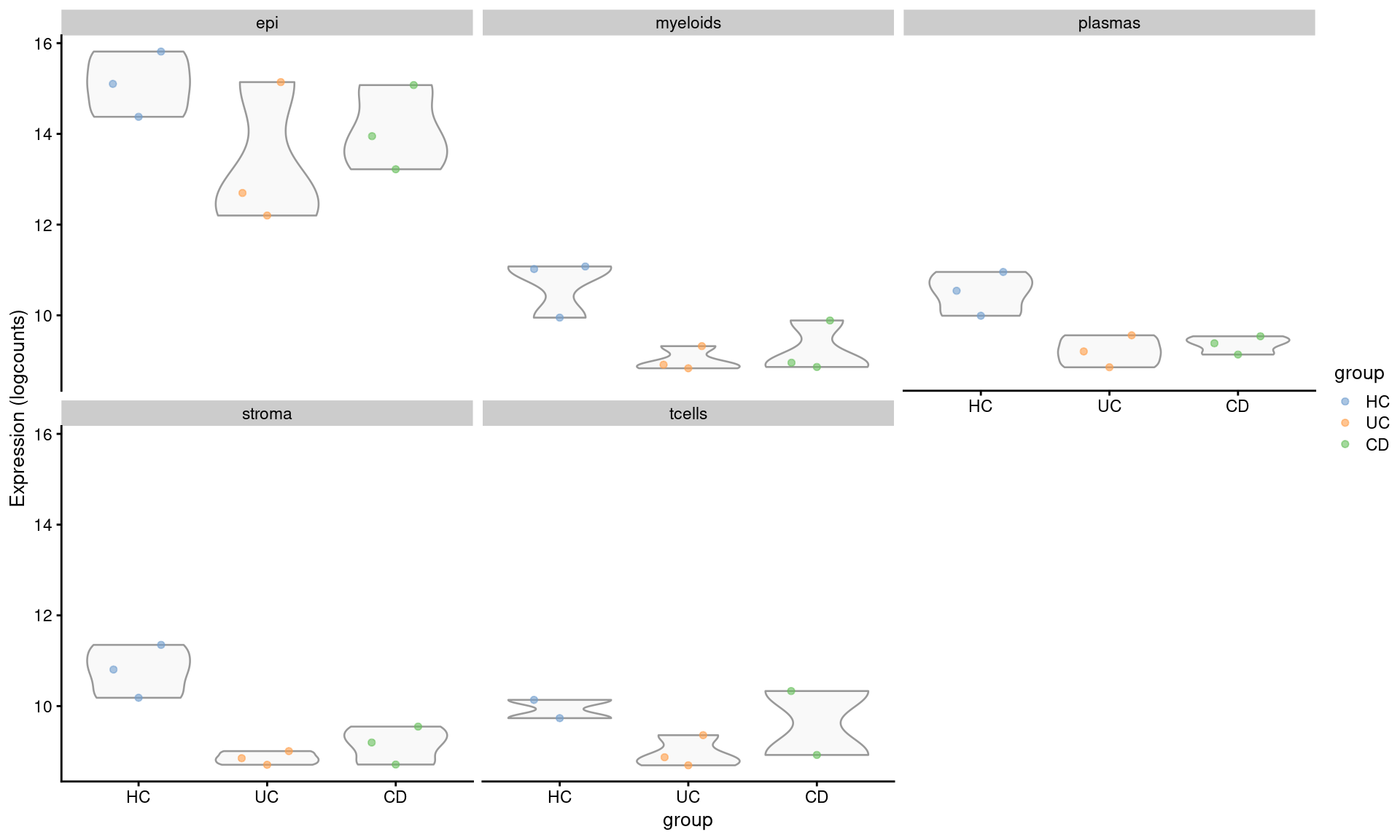
| Version | Author | Date |
|---|---|---|
| 755e4ca | swbioinf | 2025-11-07 |
Code Snippet
This code snippet would test for differential expression between groups, within each celltype or cluster independantly. It skips contrasts where there are not enough samples to test a given celltype.
It assumes the following columns in the colData cell metadata:
- total_counts : total number of counts per cell, for filtering
- celltype : Celltype grouping of cells (alternatively, cluster)
- group : Experimental group, e.g. test, control, d1_treated. In this code they are ‘GroupA’, ‘GroupB’ and ‘Control’
- tissue_sample : Unique identifier of the tissue sample cell comes from
library(SpatialFeatureExperiment)
library(scuttle)
library(tidyverse)
library(limma)
library(edgeR)
library(BiocParallel)
# CUSTOMISE:
# Set thresholds
min_reads_per_cell <- 200
min_cells_per_pdbsample <- 100
# Remove cells with too few counts
sfe <- sfe[,sfe$total_count >= min_reads_per_cell]
# CUSTOMISE:
# Define pseodoreplicate groups, with all relevant sample annotation
# remove those with too few cells.
sfe$pdb_sample <- paste0(sfe$tissue_sample,"_", sfe$celltype)
# CUSTOMISE:
## Calculate pseudobulk
#se.pdb <- aggregateAcrossCells(sfe, ids=sfe$pdb_sample)
se.pdb <- aggregateAcrossCells(sfe, ids=sfe$pdb_sample, BPPARAM = MulticoreParam(workers=8)) # Use 8 cores, requires BiocParallel.
# CUSTOMISE:
# Slow running step, highly reccomended saving pseubulk object, this is useful for plotting also.
# saveRDS(se.pdb, '~/path/to/pseudbulked_se.RDS')
# Filter on minimum number of cells per pdb group
# (ncells column was added by aggregateAcrossCells)
se.pdb <- se.pdb[,se.pdb$ncells >= min_cells_per_pdbsample]
# Pull out the pseudoboulk counts matrix for passed samples
pseudobulk_counts_matrix <- counts(se.pdb)
# And the corresponding annotations
pseudobulk_anno_table <- as_tibble(colData(se.pdb)[,c('pdb_sample','group', 'tissue_sample', 'celltype', 'ncells')]) # + any other experimental factors
# CUSTOMISE:
# Build a table of each contrast we might want to do.
#GroupAvsControl = GroupA - Control
#GroubBvsControl = GroupB - Control
contrasts_wanted <- bind_cols(
A= c("GroupA", "GroupB"), # First terms
B= c("Control", "Control") # Second terms (usually control)
)
# Empty list to collect results
de_result_list <- list()
## Cycle through each celltype
for (the_celltype in levels(se.pdb$celltype)) {
# Subset pseudobulk object to one celltype
se.pdb.this <- se.pdb[,se.pdb$celltype == the_celltype]
# And pull out the annotation and counts
anno_table.this <- as.tibble(colData(se.pdb.this))
count_matrix.this <- counts(se.pdb.this)
## Check for sufficient replicates ##
# To do any calculations, we need at least 2 pseudobulk groups per contrast.
# there are plenty in this experiemnt, but with less replicates and rare cell types
# its very common to have to skip some contrats for some celltypes.
# skip clusters with no samples after filtering
if( nrow(anno_table.this) < 1 ) {next}
# Count how many bio samples per group (only need one fov to count it)
biosample_per_group <- anno_table.this %>%
select(group, tissue_sample) %>%
pull(group) %>% # pull out group column, one entry per tissue sample
table() # count how many
# Are there enough Biological smples to consider the contrast?
min_biosample_per_group <- 2
enough_biosamples <-
(unname(biosample_per_group[contrasts_wanted$A]) >= min_biosample_per_group) &
(unname(biosample_per_group[contrasts_wanted$B]) >= min_biosample_per_group)
# Make a new table with contrasts that we have enough biosamples for.
contrasts_to_test <- contrasts_wanted[enough_biosamples,]
## Setup model
# Setup objects for limma
dge <- DGEList(count_matrix.this)
dge <- calcNormFactors(dge)
# Build model
group <- anno_table.this$group
# CUSTOMISE:
# Model design
# Add other experimental factors here
# ~0 + group
# ~0 + group + individual
# ~0 + group + individual + slide
design <- model.matrix( ~0 + group)
# Run Voom
vm <- voom(dge, design = design, plot = FALSE)
# Adding dupliate correlation to use individual fovs, rather than pooled per biosample
fit <- lmFit(vm, design)
# Define and fit contrasts and run ebayes
# Doing this in an automated way so we can include/exvlude contrasts where
# there are / arenot enough replicates available.
# That's why generatgin a string .
contrast_str_list <- paste0("group",contrasts_to_test$A,"-","group",contrasts_to_test$B)
contrasts <- makeContrasts(contrasts=contrast_str_list,
levels=coef(fit))
fit <- contrasts.fit(fit, contrasts)
fit <- eBayes(fit)
## Loop through contrasts.
# You ccan run multiple contrasts at onces, but doing it this way allows us
# to skip individual contrasts within a celltype.
for ( the_coef in colnames(contrasts) ) {
# Make a version of the coeffient name that doesn't include a '-', or the
# factor name (group)
# Life is easier without special characters
# groupUC-groupHC => UCvHC
contrast_name <- gsub("group","", gsub("-","v",the_coef))
de_result.this <- topTable(fit, n = Inf, adjust.method = "BH", coef = the_coef) %>%
rownames_to_column("target") %>%
mutate(contrast=contrast_name,
contrast_group="pairwise",
celltype=the_celltype) %>%
select(celltype,contrast_group, contrast,target,everything()) %>%
arrange(P.Value)
# build a unique name for this result by adding
# <celltype>_<A>v<B>
de_result_list[[paste(the_celltype, contrast_name, sep="_")]] <- de_result.this
}
}
# Join together results for all celltypes, and pull out those with a singificant adjusted p-value
de_results_all <- bind_rows(de_result_list)
de_results_sig <- filter(de_results_all, adj.P.Val < 0.05)Results
DT::datatable(mutate(head(de_results_sig), across(is.numeric, signif, digits = 3)))This table is the typical output of limma tests; With a couple of extra columns added by our code.
- celltype: The celltype being tested (Added by example code)
- contrast: The contrast being tested (Added by example code)
- target : The gene name (Added by example code, is the rowname in limma output)
- rownames : The tested cell types
- logFC : Log 2 fold change between tested groups.
For a test of A-B;
- At logFC +1, A is doubled B.
- At logFC -1, A is half of B.
- A logFC 0 indicates no change.
- AveExpr : Average expression of a gene across all replicates.
- t : Moderated T-statistic. See Limma documentation.
- P.Value : P.value
- adj.P.Val : A multiple-hypothesis corrected p-value
- B : B statistic (rarely used). See Limma documentation.
More Information
- ‘Ochestrating single cell analysis with bioconductor’ book chapter ‘DE analyses between conditions’ : An explanation of the ‘pseudobulk’ approch to single cell differential expression calculation.
- Pseudobulk and differential expression (glmGamPoi documentation): Part of the documentation for the glmGamPoi R package (not used here), that features another clear explanation of pseudobulking.
- Bias, robustness and scalability in single-cell differential expression analysis (Soneson and Robinson 2018) : A review of single cell differential expression calculation methods.
- limma documentation (Ritchie et al. 2015): The complete manual to limma.
- Use of within-array replicate spots for assessing differential expression in microarray experiments(Smyth, Michaud, and Scott 2005) : Paper describing the duplicate correlation approach for pseudoreplicates (in the original context of micorarray analysis).
- Bioconductor support: Further clarification on when not to use duplicateCorrelation with technical replicates (RNA-seq) : A very helpful discussion about when to use correlateDuplicates for pseuodreplicates.
- Differential Expression with Limma-Voom UC davis bioinformatics training : A more accessible explanation of bulk RNAseq analyses using limma.
- Interactions and contrasts : An excellent visual explanation of how to build linear models for more complex multi-factor experimental designs (e.g. treatment and genotype). Part of a larger Data Analysis for Genomics class resource.
- Voyager Website: Voyager is a great package for exploring and visualising spatial data around the ‘SpatialFeatureExperiment’ (SFE) format. Many of the useful plots are from this package.
References
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 22.04.5 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.20.so; LAPACK version 3.10.0
locale:
[1] LC_CTYPE=en_AU.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_AU.UTF-8 LC_COLLATE=en_AU.UTF-8
[5] LC_MONETARY=en_AU.UTF-8 LC_MESSAGES=en_AU.UTF-8
[7] LC_PAPER=en_AU.UTF-8 LC_NAME=en_AU.UTF-8
[9] LC_ADDRESS=en_AU.UTF-8 LC_TELEPHONE=en_AU.UTF-8
[11] LC_MEASUREMENT=en_AU.UTF-8 LC_IDENTIFICATION=en_AU.UTF-8
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats4 stats graphics grDevices datasets utils methods
[8] base
other attached packages:
[1] ggrepel_0.9.6 patchwork_1.3.2
[3] Voyager_1.13.0 BiocParallel_1.44.0
[5] edgeR_4.8.1 DT_0.34.0
[7] limma_3.66.0 lubridate_1.9.4
[9] forcats_1.0.1 stringr_1.6.0
[11] dplyr_1.1.4 purrr_1.2.0
[13] readr_2.1.6 tidyr_1.3.1
[15] tibble_3.3.0 tidyverse_2.0.0
[17] scater_1.38.0 ggplot2_4.0.1
[19] scuttle_1.20.0 SpatialExperiment_1.20.0
[21] SingleCellExperiment_1.32.0 SummarizedExperiment_1.40.0
[23] Biobase_2.70.0 GenomicRanges_1.62.1
[25] Seqinfo_1.0.0 IRanges_2.44.0
[27] S4Vectors_0.48.0 BiocGenerics_0.56.0
[29] generics_0.1.4 MatrixGenerics_1.22.0
[31] matrixStats_1.5.0 alabaster.sfe_1.2.0
[33] alabaster.base_1.10.0 SpatialFeatureExperiment_1.12.1
[35] workflowr_1.7.2
loaded via a namespace (and not attached):
[1] fs_1.6.6 spatialreg_1.4-2
[3] bitops_1.0-9 sf_1.0-23
[5] EBImage_4.52.0 httr_1.4.7
[7] RColorBrewer_1.1-3 tools_4.5.2
[9] utf8_1.2.6 R6_2.6.1
[11] HDF5Array_1.38.0 rhdf5filters_1.22.0
[13] withr_3.0.2 sp_2.2-0
[15] gridExtra_2.3 cli_3.6.5
[17] RBioFormats_1.10.0 sandwich_3.1-1
[19] alabaster.se_1.10.0 labeling_0.4.3
[21] sass_0.4.10 mvtnorm_1.3-3
[23] arrow_22.0.0 S7_0.2.1
[25] proxy_0.4-28 R.utils_2.13.0
[27] dichromat_2.0-0.1 scico_1.5.0
[29] rstudioapi_0.17.1 crosstalk_1.2.2
[31] spdep_1.4-1 Matrix_1.7-4
[33] ggbeeswarm_0.7.3 abind_1.4-8
[35] R.methodsS3_1.8.2 terra_1.8-86
[37] lifecycle_1.0.4 whisker_0.4.1
[39] multcomp_1.4-29 yaml_2.3.12
[41] rhdf5_2.54.1 SparseArray_1.10.7
[43] grid_4.5.2 promises_1.5.0
[45] dqrng_0.4.1 alabaster.spatial_1.10.0
[47] lattice_0.22-7 cowplot_1.2.0
[49] beachmat_2.26.0 magick_2.9.0
[51] zeallot_0.2.0 pillar_1.11.1
[53] knitr_1.50 rjson_0.2.23
[55] boot_1.3-32 sfarrow_0.4.1
[57] codetools_0.2-20 wk_0.9.4
[59] glue_1.8.0 getPass_0.2-4
[61] data.table_1.17.8 memuse_4.2-3
[63] vctrs_0.6.5 png_0.1-8
[65] gtable_0.3.6 assertthat_0.2.1
[67] cachem_1.1.0 xfun_0.55
[69] S4Arrays_1.10.1 DropletUtils_1.30.0
[71] coda_0.19-4.1 survival_3.8-3
[73] sfheaders_0.4.5 rJava_1.0-11
[75] units_1.0-0 statmod_1.5.1
[77] bluster_1.20.0 TH.data_1.1-5
[79] nlme_3.1-168 bit64_4.6.0-1
[81] alabaster.ranges_1.10.0 BumpyMatrix_1.18.0
[83] rprojroot_2.1.1 bslib_0.9.0
[85] irlba_2.3.5.1 vipor_0.4.7
[87] KernSmooth_2.23-26 otel_0.2.0
[89] spData_2.3.4 DBI_1.2.3
[91] tidyselect_1.2.1 processx_3.8.6
[93] bit_4.6.0 compiler_4.5.2
[95] git2r_0.36.2 BiocNeighbors_2.4.0
[97] h5mread_1.2.1 xml2_1.5.1
[99] DelayedArray_0.36.0 scales_1.4.0
[101] classInt_0.4-11 callr_3.7.6
[103] tiff_0.1-12 digest_0.6.39
[105] fftwtools_0.9-11 alabaster.matrix_1.10.0
[107] rmarkdown_2.30 XVector_0.50.0
[109] htmltools_0.5.9 pkgconfig_2.0.3
[111] jpeg_0.1-11 sparseMatrixStats_1.22.0
[113] fastmap_1.2.0 rlang_1.1.6
[115] htmlwidgets_1.6.4 DelayedMatrixStats_1.32.0
[117] farver_2.1.2 jquerylib_0.1.4
[119] zoo_1.8-15 jsonlite_2.0.0
[121] R.oo_1.27.1 BiocSingular_1.26.1
[123] RCurl_1.98-1.17 magrittr_2.0.4
[125] s2_1.1.9 Rhdf5lib_1.32.0
[127] Rcpp_1.1.0 ggnewscale_0.5.2
[129] viridis_0.6.5 stringi_1.8.7
[131] alabaster.schemas_1.10.0 MASS_7.3-65
[133] alabaster.bumpy_1.10.0 parallel_4.5.2
[135] deldir_2.0-4 splines_4.5.2
[137] hms_1.1.4 locfit_1.5-9.12
[139] ps_1.9.1 igraph_2.2.1
[141] ScaledMatrix_1.18.0 LearnBayes_2.15.1
[143] evaluate_1.0.5 renv_1.1.5
[145] BiocManager_1.30.27 tzdb_0.5.0
[147] httpuv_1.6.16 alabaster.sce_1.10.0
[149] rsvd_1.0.5 e1071_1.7-16
[151] RSpectra_0.16-2 later_1.4.4
[153] viridisLite_0.4.2 class_7.3-23
[155] beeswarm_0.4.0 cluster_2.1.8.1
[157] timechange_0.3.0